Blob Columns¶
Lance supports large binary objects (images, videos, audio, model artifacts) through blob columns.
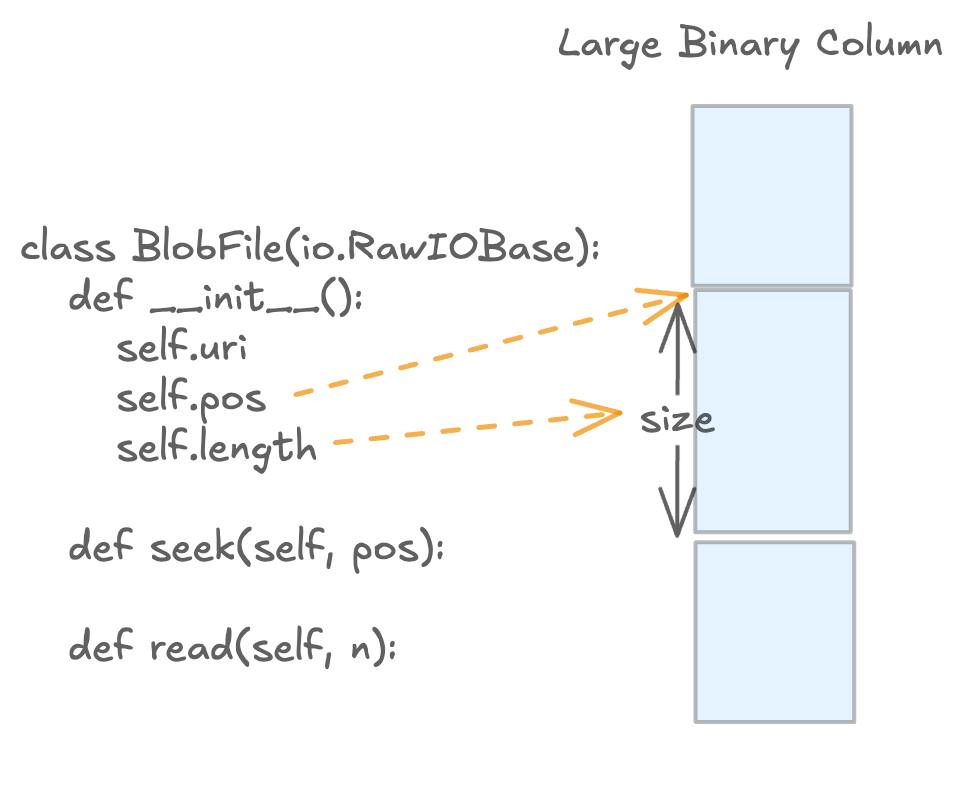
Blob access is lazy: reads return BlobFile handles so callers can stream bytes on demand.
What This Page Covers¶
This page focuses on Python blob workflows and uses Lance file format terminology.
data_storage_versionmeans the Lance file format version of a dataset.- A dataset's
data_storage_versionis fixed once the dataset is created. - If you need a different file format version, write a new dataset.
Quick Start (Blob v2)¶
import lance
import pyarrow as pa
from lance import blob_array, blob_field
schema = pa.schema([
pa.field("id", pa.int64()),
blob_field("blob"),
])
table = pa.table(
{
"id": [1],
"blob": blob_array([b"hello blob v2"]),
},
schema=schema,
)
ds = lance.write_dataset(table, "./blobs_v22.lance", data_storage_version="2.2")
blob = ds.take_blobs("blob", indices=[0])[0]
with blob as f:
assert f.read() == b"hello blob v2"
Version Compatibility (Single Source of Truth)¶
Dataset data_storage_version |
Legacy blob metadata (lance-encoding:blob) |
Blob v2 (lance.blob.v2) |
|---|---|---|
0.1, 2.0, 2.1 |
Supported for write/read | Not supported |
2.2+ |
Not supported for write | Supported for write/read (recommended) |
Important:
- For file format
>= 2.2, legacy blob metadata (lance-encoding:blob) is rejected on write.
Blob v2 Write Patterns¶
Use blob_field and blob_array to build blob v2 columns.
import lance
import pyarrow as pa
from lance import Blob, blob_array, blob_field
schema = pa.schema([
pa.field("id", pa.int64()),
blob_field("blob", nullable=True),
])
# A single column can mix:
# - inline bytes
# - external URI
# - external URI slice (position + size)
# - null
rows = pa.table(
{
"id": [1, 2, 3, 4],
"blob": blob_array([
b"inline-bytes",
"s3://bucket/path/video.mp4",
Blob.from_uri("s3://bucket/archive.tar", position=4096, size=8192),
None,
]),
},
schema=schema,
)
ds = lance.write_dataset(
rows,
"./blobs_v22.lance",
data_storage_version="2.2",
)
Note:
- By default, external blob URIs must map to a registered non-dataset-root base path.
- If you need to reference external objects outside those bases, set
allow_external_blob_outside_bases=Truewhen writing.
Example: packed external blobs (single container file)¶
import io
import tarfile
from pathlib import Path
import lance
import pyarrow as pa
from lance import Blob, blob_array, blob_field
# Build a tar file with three payloads
payloads = {
"a.bin": b"alpha",
"b.bin": b"bravo",
"c.bin": b"charlie",
}
with tarfile.open("container.tar", "w") as tf:
for name, data in payloads.items():
info = tarfile.TarInfo(name)
info.size = len(data)
tf.addfile(info, io.BytesIO(data))
# Capture offset/size for each member
blob_values = []
with tarfile.open("container.tar", "r") as tf:
container_uri = Path("container.tar").resolve().as_uri()
for name in payloads:
m = tf.getmember(name)
blob_values.append(Blob.from_uri(container_uri, position=m.offset_data, size=m.size))
schema = pa.schema([
pa.field("name", pa.utf8()),
blob_field("blob"),
])
rows = pa.table(
{
"name": list(payloads.keys()),
"blob": blob_array(blob_values),
},
schema=schema,
)
ds = lance.write_dataset(
rows,
"./packed_blobs_v22.lance",
data_storage_version="2.2",
allow_external_blob_outside_bases=True,
)
Blob v2 Read Patterns¶
Use take_blobs to fetch file-like handles.
Exactly one selector must be provided: ids, indices, or addresses.
| Selector | Typical Use | Stability |
|---|---|---|
indices |
Positional reads within one dataset snapshot | Stable within that snapshot |
ids |
Logical row-id based reads | Stable logical identity (when row ids are available) |
addresses |
Low-level physical reads and debugging | Unstable physical location |
Read by row indices¶
import lance
ds = lance.dataset("./blobs_v22.lance")
blobs = ds.take_blobs("blob", indices=[0, 1])
with blobs[0] as f:
data = f.read()
Read by row ids¶
import lance
ds = lance.dataset("./blobs_v22.lance")
row_ids = ds.to_table(columns=[], with_row_id=True).column("_rowid").to_pylist()
blobs = ds.take_blobs("blob", ids=row_ids[:2])
Read by row addresses¶
import lance
ds = lance.dataset("./blobs_v22.lance")
row_addrs = ds.to_table(columns=[], with_row_address=True).column("_rowaddr").to_pylist()
blobs = ds.take_blobs("blob", addresses=row_addrs[:2])
Example: decode video frames lazily¶
import av
import lance
ds = lance.dataset("./videos_v22.lance")
blob = ds.take_blobs("video", indices=[0])[0]
start_ms, end_ms = 500, 1000
with av.open(blob) as container:
stream = container.streams.video[0]
stream.codec_context.skip_frame = "NONKEY"
start = (start_ms / 1000) / stream.time_base
end = (end_ms / 1000) / stream.time_base
container.seek(int(start), stream=stream)
for frame in container.decode(stream):
if frame.time is not None and frame.time > end_ms / 1000:
break
# process frame
pass
Legacy Compatibility Appendix (data_storage_version <= 2.1)¶
If you need to keep writing legacy blob columns, use file format 0.1, 2.0, or 2.1
and mark LargeBinary fields with lance-encoding:blob = true.
import lance
import pyarrow as pa
schema = pa.schema([
pa.field("id", pa.int64()),
pa.field(
"video",
pa.large_binary(),
metadata={"lance-encoding:blob": "true"},
),
])
table = pa.table(
{
"id": [1, 2],
"video": [b"foo", b"bar"],
},
schema=schema,
)
ds = lance.write_dataset(
table,
"./legacy_blob_dataset",
data_storage_version="2.1",
)
This write pattern is invalid for data_storage_version >= 2.2.
For new datasets, prefer blob v2.
Rewrite to a New Blob v2 Dataset¶
If your current dataset is legacy blob and you want blob v2, rewrite into a new dataset with data_storage_version="2.2".
import lance
import pyarrow as pa
from lance import blob_array, blob_field
legacy = lance.dataset("./legacy_blob_dataset")
raw = legacy.scanner(columns=["id", "video"], blob_handling="all_binary").to_table()
new_schema = pa.schema([
pa.field("id", pa.int64()),
blob_field("video"),
])
rewritten = pa.table(
{
"id": raw.column("id"),
"video": blob_array(raw.column("video").to_pylist()),
},
schema=new_schema,
)
lance.write_dataset(
rewritten,
"./blob_v22_dataset",
data_storage_version="2.2",
)
Warning:
- The example above materializes binary payloads in memory (
blob_handling="all_binary"andto_pylist()). - For large datasets, prefer chunked/batched rewrite pipelines.
Troubleshooting¶
"Blob v2 requires file version >= 2.2"¶
Cause:
- You are writing blob v2 values into a dataset/file format below
2.2.
Fix:
- Write to a dataset created with
data_storage_version="2.2"(or newer).
"Legacy blob columns ... are not supported for file version >= 2.2"¶
Cause:
- You are using legacy blob metadata (
lance-encoding:blob) while writing2.2+data.
Fix:
- Replace legacy metadata-based columns with blob v2 columns (
blob_field/blob_array).
"Exactly one of ids, indices, or addresses must be specified"¶
Cause:
take_blobsreceived none or multiple selectors.
Fix:
- Provide exactly one of
ids,indices, oraddresses.